An ongoing question regarding the SARS-CoV-2 virus is the extent of variation. The Omicron family of viruses are among the most mutated natural variants observed thus far, displaying over fifty amino acid changes throughout the genome. While some may hope that this approaches an upper limit, the unfortunate truth is that the potential number of mutations is unknowable due to the potential of animal cross-infection. Here we describe a recent paper by Pickering et al. that describes a series of SARS-CoV-2 sequences originating in white-tailed deer highly divergent from previous variants, with as many as 76 consensus nucleotide mutations from the original Wuhan virus.
Instances of SARS-CoV-2 infecting animals and yielding new variants is not uncommon. We have observed emerging variant strains from Gorillas in California zoos with Epsilon and mink in Denmark and the United States. These variants reflect adaptations to the new host species. This reflects documented transmission from hamsters, mink, and deer into humans.
Several previous studies have documented SARS-CoV-2 infection of white-tailed deer. Palmer et al. describe the isolation of multiple independent isolates of SARS-CoV-2 in US deer populations. The sequence of these isolates is very close or identical to that of prevailing human isolates in the same geographical area. That renders the discovery of a new highly-diverged variant in Canadian deer a troubling outlier.
Now the story from Ontario, Canada. White-tailed deer regularly come in contact with humans through hunting and foraging. This species is among the most common large mammal in North America and regularly contact humans via hunting activity. Pickering et al. collected SARS-CoV-2 samples from white-tailed deer to examine new mutations and monitor for similar associated human cases, i.e., deer-to-human transmission. Of over 250 sampled deer, roughly 6% had antibodies in the sera samples to SARS-CoV-2, indicating previous infection.
From five nasal swabs taken from white-tailed deer infected with SARS-CoV-2, Pickering et al. developed three consensus genomes that contained 76 conserved nucleotide mutations. In searching for the deer-to-human transmission, they also found a SARS-CoV-2 genome in the GISAID database derived from a human host in Michigan in November 2020 that closely resembles the deer consensus.
The surprise comes with the striking difference between the Ontario white-tailed deer samples and previous variants of concern or interest. Figure 1 illustrates the departure of the 76 consensus deer mutations as compared to other animal derived sequences in the GISAID database.
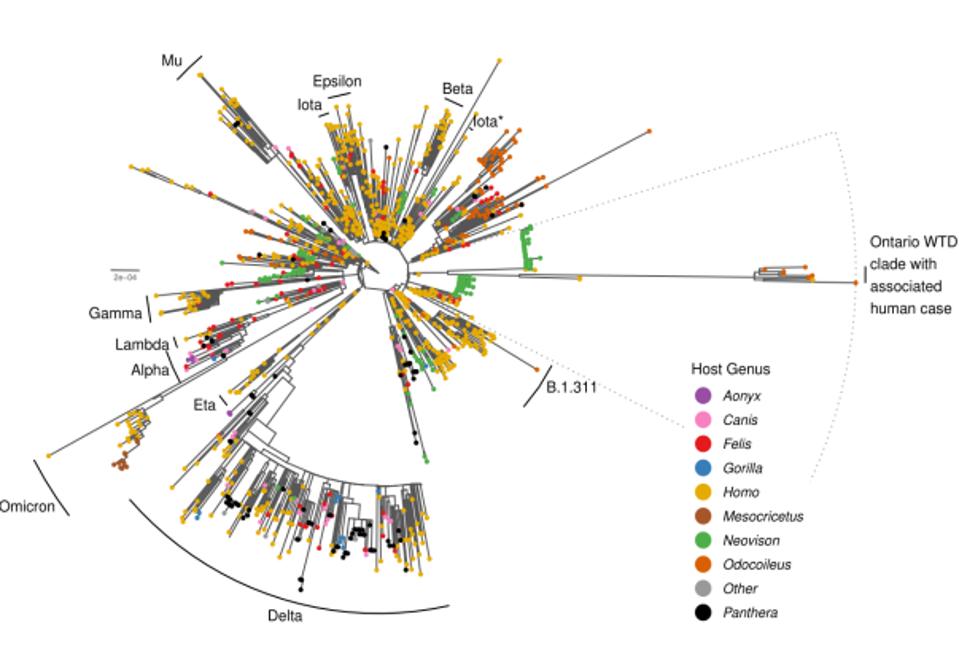
In tracking down the origin of this strain, the authors note it likely stemmed in late 2020 from Beta, Iota, or Epsilon variant. These variants were also circulating mink populations in Wisconsin and Michigan in late 2020. Pickering et al. believe that at some point thereafter, the strain transmitted into the Ontario deer population via an intermediary unknown species. Pickering et al. note that the large number of mutations specifically occurred in the undetected intermediary for a prolonged period, up until their detection in this study. They entertain the notion that the intermediary may have been humans. Based on the sheer number of species that interact with humans regularly, there are likely several reverse zoonosis strains akin to the white-tailed deer strain waiting to emerge. The potential severity of the strains is uncertain.
As outlined in Figure 2, after a long residence in the unknown predecessor, the white-tailed deer strain either made its way into deer and the one human concurrently or sequentially. This gives us some insight into the blueprint of reverse zoonosis strains.
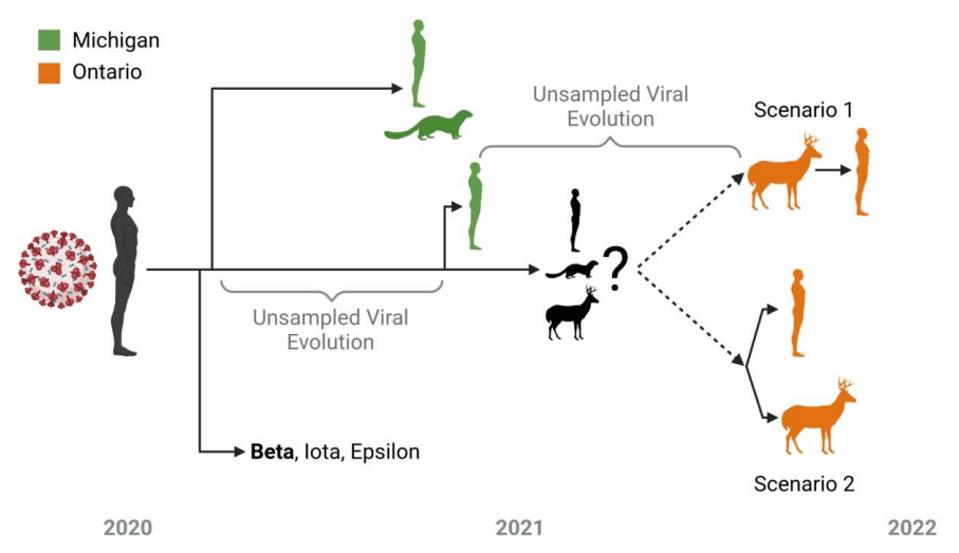
We note that this series of events resembles, in some ways, the origin of the Omicron family of viruses (Figure 5). From the original Wuhan strain sampled in humans in late 2019, the B.1 virus which we call the Triad emerged in February and March 2020. This variant contained a triad of mutations: D614G in the Spike, P323L in NSP12, and C241U in the 5’ UTR. We previously proposed that an undetected parent of the Omicron viruses emerged from B.1 containing 31 mutations. From there, at least five distinct strains have been sampled in recent months, comprising the Omicron family.
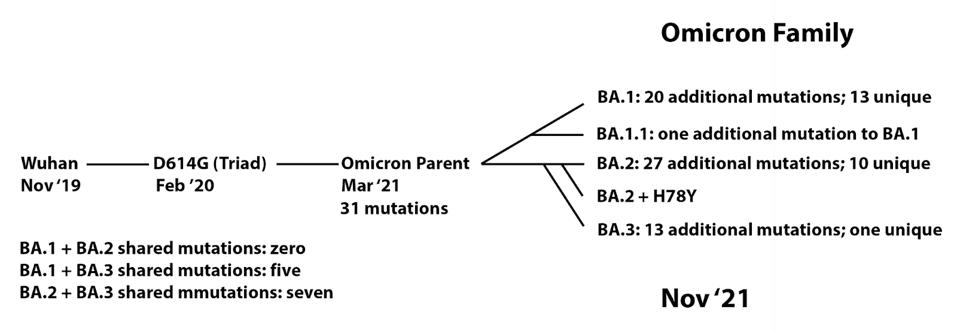
Of the 76 conserved mutations, Pickering et al. observed that many were not found in human or mink samples in the region at the time of collection, aside from the one human sample taken from Michigan months prior.
Of the 76 nucleotide changes, 49 are different from the closest related natural variant. There are 51 changes in Orf1ab, 11 of which are in NSP3 and nine in NSP4. There are nine changes in the Spike protein. The structural proteins E, M, and N, contain six observed mutations and the Orf genes contain ten. Below is a diagram of the sequenced mutations in the deer consensus and human genomes. Additionally, the deer strains have the C241U conserved mutation in the 5’ untranslated region of the genome associated with cellular TAR binding.
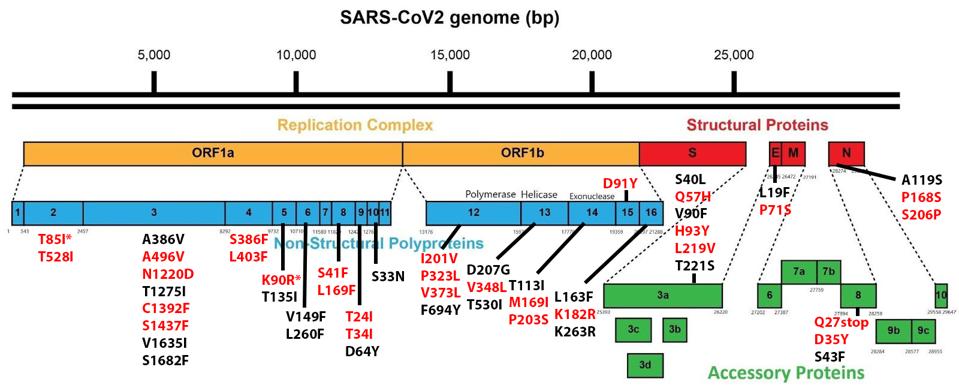
The mutations noted, however, were not exceptionally uncommon in animal sequences. Particularly in the Spike protein, which interacts with the human or nonhuman ACE2 receptor, the deer variant contains many mutations shared with bats, cats, hamsters, and mink. Of the consensus Spike protein amino acid mutations (H49Y, T95I, 143-145del, F486L, N501T, and D614G), only F486L is rarely found in any sequence, human or other. There were also three amino acid mutations in only one white-tailed deer genome (T22I, S247G, and V705A) and several cis-acting synonymous mutations that do not change amino acids but could affect RNA transcription sequences.
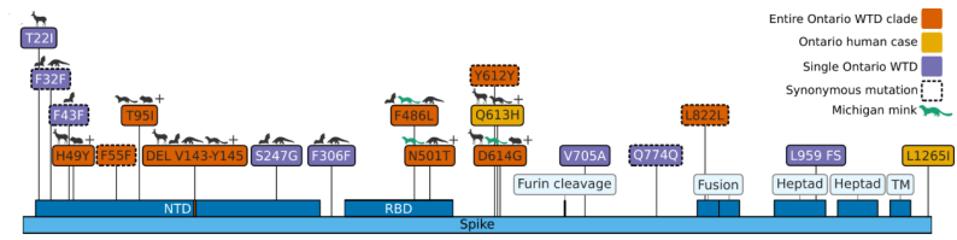
Note that most mutations in the strain are not in the Spike protein. The majority are in the Orf1ab replicative proteins, structural proteins, and accessory genes. Pickering et al. make the unusual suggestion that the majority of these mutations arise without selection, simply via genetic drift. The difference in mutations between the deer samples and Omicron, particularly in the Spike protein, illustrate the differences in selective pressure, as the paucity of mutations in the deer Spike likely indicate less efficient transmissibility as compared to Omicron. The large number of mutations outside of the Spike may represent either neutral drift or other selective pressures, including T-cell recognition and suppression of innate immune responses.
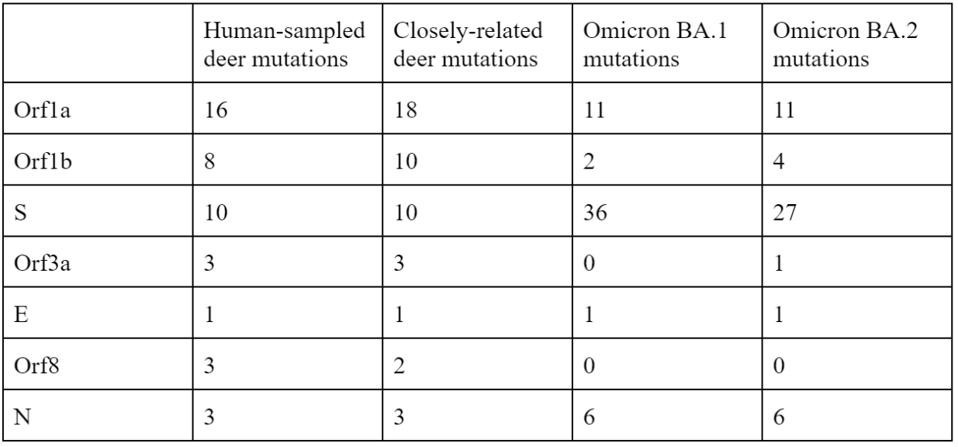
The relative lack of mutations in the Spike protein is also reflected by the comparable neutralization of these viruses by vaccine-elicited antibodies to non-Omicron variants. Additionally, it does not seem that infected deer were significantly affected by symptoms, similar to the Omicron variant in reduced symptom severity concentrated in the nasopharynx.
This observation should put us on notice that there may be many other lingering variants in humans or in other species, which at any time may reemerge to cause the kind of pandemic associated with Omicron. Instances of reverse zoonosis are more common than we realize, as many households have domesticated pets and many cities have many wild and feral species. As a final note, hunters in this region should be wary of dressing deer carcasses as this represents a transmission risk.








